Emails from eBay members asking questions come through as randomstring@members.ebay.com which can make them hard to count as a group
The following R script re-codes them so they can be counted.
install.packages("plyr")
library(plyr)
setwd("/home/mike/Desktop")
dir()
inbox<-read.csv("inbox.TXT", sep="\t")
names(inbox)
inbox$EADD <- ifelse(grepl("members.ebay.co.uk",inbox$From...Address.),
"members.ebay.co.uk" ,
c(as.character(inbox$From...Address.)))
str(inbox)
f <- ddply(inbox,c("EADD"),summarize,N=length(EADD))
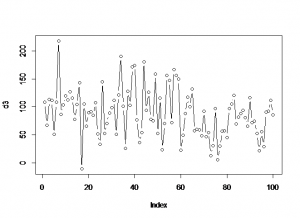
plot(f$N)
head(f[order(-f$N),])
str(f)
Once you've identified the biggest culprits, make a rule to move or delete them from your email.