Read the clipboard into R:
DataFrame<-read.table("clipboard",headers=TRUE)
Write out the data in a Dataframe into the clipboard:
write.table(DataFrame,"clipboard-2048",sep="\t")
You can use this to paste data into other applications.
Create a DataFrame filled with random data (there may be a shorter way of doing this)
x1<-rnorm(2000,750,55)
x2 <-rnorm(2000,16,100)
df1 <-data.frame(cbind("D",x1,x2))
Here we've used "cbind" which means bind some columns
the data.frame bit seems to convert something into a collection of columns into a DataFrame.
x1<-rnorm(2000,500,155)
x2<-rnorm(2000,250,100)
df2<-data.frame(cbind("A",x1,x2))
Now join them together...
df0<-data.frame(rbind(df1,df2))
Now for some reason we have to convert from factors to numerics...
df0$X1<-as.numeric(as.character(df0$x1))
df0$X2<-as.numeric(as.character(df0$x2))
Notice that we've added two extra columns x1 has been converted into X1 and x2 to X2
str(df0)
Finally let's plot them:
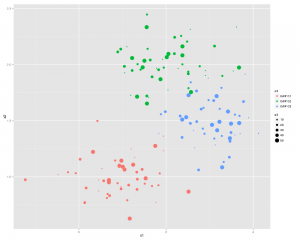
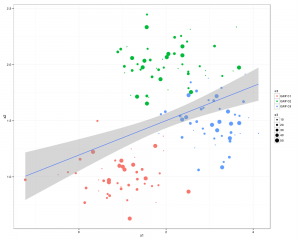
scatterplot(X1 ~X2 | V1 ,data=df0,smooth=FALSE,ellipse=FALSE,lty=0)
Which will look something like this:

We can also generate a boxplot using:
bwplot(X2~V1,data = df0)
boxplot(X2~V1,data = df0)
bwplot requires the lattice library.

Back to the scatter plot graph:
scatterplot(X1 ~X2 | V1
,data=df0
,smooth=FALSE
,ellipse=FALSE
,lty=0
,grid=TRUE
,boxplots="xy")
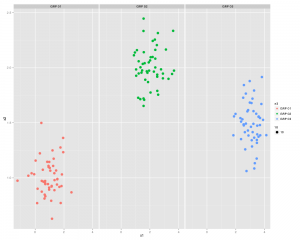
Which gives something like this:

Here the boxplots are describing all of X1 and X2 i.e. X1 of A and D combined.